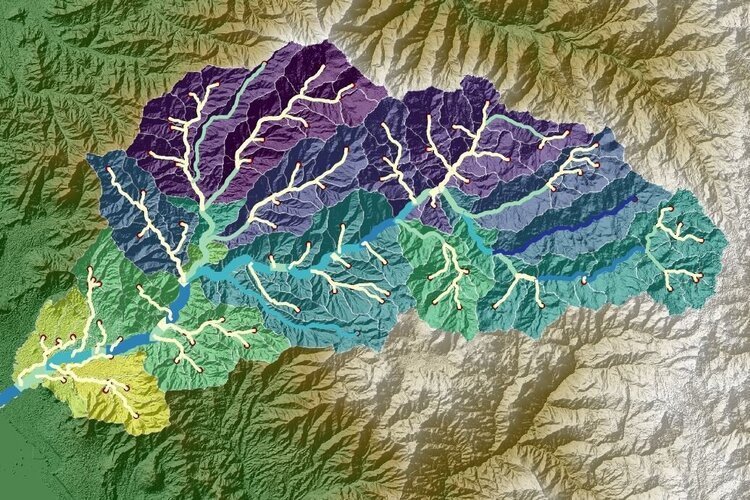
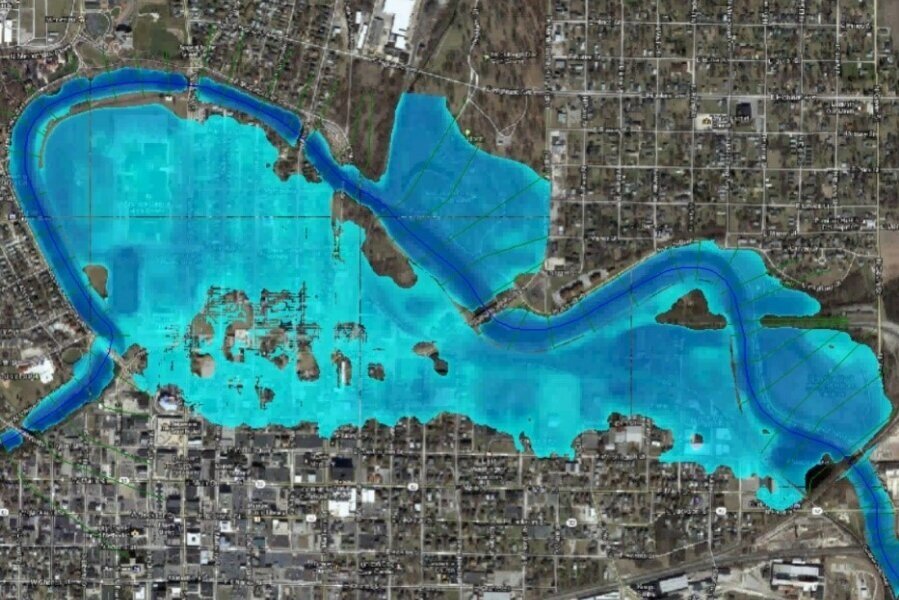
How to create a geospatial Raster from XY data with Python, Pandas and Rasterio - Tutorial
/Another tutorial done under the concept of “geospatial python”. The tutorial shows the procedure to run a Scipy interpolation over a Pandas dataframe of point related data having a 2D Numpy array as an output. With some procedures of Rasterio the Numpy array was transformed into a monoband geospatial Tiff raster.
Tutorial
Enjoy the videos and music you love, upload original content, and share it all with friends, family, and the world on YouTube.
Code
import numpy as np
import pandas as pd
from scipy.interpolate import griddata
import matplotlib.pyplot as plt
import rasterio#open the chemistry data
chemData = pd.read_csv('../tab/chemistryData.csv',index_col=0,usecols=[0,5,6,18])
chemData = chemData.dropna() #delete missing data rows
chemData.describe()| EsteCor | NorteCor | pH | |
|---|---|---|---|
| count | 176.000000 | 1.760000e+02 | 176.000000 |
| mean | 247362.431818 | 8.352024e+06 | 8.070892 |
| std | 5246.316476 | 7.225883e+03 | 0.877716 |
| min | 236536.000000 | 8.336049e+06 | 5.630000 |
| 25% | 242377.750000 | 8.346304e+06 | 7.617500 |
| 50% | 249067.000000 | 8.352004e+06 | 8.045000 |
| 75% | 251132.000000 | 8.356741e+06 | 8.510000 |
| max | 259877.000000 | 8.372586e+06 | 10.400000 |
# Optional : drop extreme values
chemData = chemData[chemData.pH > np.quantile(chemData.pH,0.1)]
chemData = chemData[chemData.pH < np.quantile(chemData.pH,0.9)]
chemData.describe()| EsteCor | NorteCor | pH | |
|---|---|---|---|
| count | 140.000000 | 1.400000e+02 | 140.000000 |
| mean | 247838.157143 | 8.352633e+06 | 8.076693 |
| std | 5230.177137 | 7.042238e+03 | 0.489108 |
| min | 236536.000000 | 8.336049e+06 | 7.100000 |
| 25% | 243042.750000 | 8.347914e+06 | 7.790000 |
| 50% | 250146.000000 | 8.353390e+06 | 8.055000 |
| 75% | 251441.250000 | 8.357699e+06 | 8.370000 |
| max | 259877.000000 | 8.372586e+06 | 9.330000 |
#define interpolation inputs
points = list(zip(chemData.EsteCor,chemData.NorteCor))
values = chemData.pH.values
print(points[:5])
print(values[:5])[(250901.0, 8342712.0), (244329.0, 8346119.0), (244329.0, 8346119.0), (244329.0, 8346119.0), (240476.0, 8351811.000000001)]
[8.03 8.96 7.8 8.04 8.17]#define raster resolution
rRes = 50
#create coord ranges over the desired raster extension
xRange = np.arange(chemData.EsteCor.min(),chemData.EsteCor.max()+rRes,rRes)
yRange = np.arange(chemData.NorteCor.min(),chemData.NorteCor.max()+rRes,rRes)
print(xRange[:5],yRange[:5])[236536. 236586. 236636. 236686. 236736.] [8336049. 8336099. 8336149. 8336199. 8336249.]#create arrays of x,y over the raster extension
gridX,gridY = np.meshgrid(xRange, yRange)
#interpolate over the grid
gridPh = griddata(points, values, (gridX,gridY), method='linear')#show interpolated values
plt.imshow(gridPh)#definition of the raster transform array
from rasterio.transform import Affine
transform = Affine.translation(gridX[0][0]-rRes/2, gridY[0][0]-rRes/2)*Affine.scale(rRes,rRes)
transformAffine(50.0, 0.0, 236511.0,
0.0, 50.0, 8336023.999999999)#get crs as wkt
from rasterio.crs import CRS
rasterCrs = CRS.from_epsg(32718)
rasterCrs.data{'init': 'epsg:32718'}#definition, register and close of interpolated raster
interpRaster = rasterio.open('../rst/interpRaster3.tif',
'w',
driver='GTiff',
height=gridPh.shape[0],
width=gridPh.shape[1],
count=1,
dtype=gridPh.dtype,
crs=rasterCrs,
transform=transform,
)
interpRaster.write(gridPh,1)
interpRaster.close()Input data
You can download the input data from this link.